Genetic Fingerprinting at the Wisconsin Cranberry Research Station: Part 2
After the cultivars present in each bed at the newly-dedicated Wisconsin Cranberry Research Station were fingerprinted and named (see CCMJ 35.2), Dr. Zalapa’s lab was able to compare historic yields with the purity of the vines in each bed. They were also able to compare the visual assessment (height, color, and yielding/barren) of cultivar contamination, with the measured genetic contamination.
For yield comparisons, the Stevens beds had yield data available for 2013, 2014, and 2017; while the BG bed was more recently planted and only had 2017 yield data. When viewing the table, remember the beds were numbered 1 to 10, with the first bed being furthest south and bed 10 being the furthest north. Contamination in beds ranged from 0% to 69%, although yield did not track percent contamination as strongly as we had suspected. Figure 1 displays how yield reduces with % contamination—there is a negative trend but it only describes SOME of the yield variation. You can see that some beds did much better than their contamination % would have suggested (ie bed 3), while other beds did more poorly than expected (ie bed 2). Chart 2 lists the raw data for each bed.
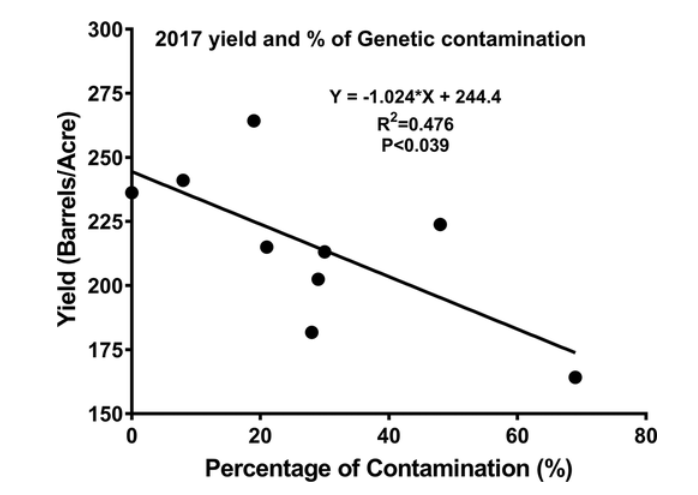
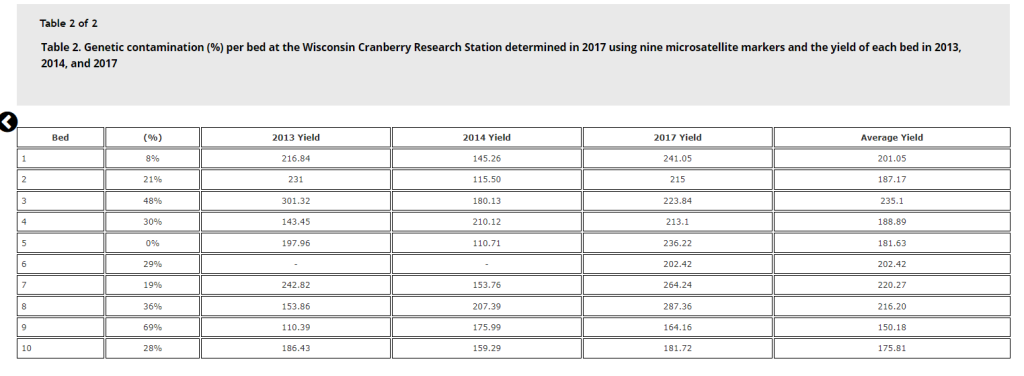
In comparing visual determination of cultivar from genetic determination, in general visual detection was able to distinguish Perry Red contaminants from Stevens vines. It appears that the Perry Red contamination was planted in alternating fashion, possibly due to the mowing and baling practice used for planting from genetically untested cuttings.
There were some instances in which the visual examination noticed “differences” which were not there—the researcher thought they were viewing Perry Red, but in fact were seeing Stevens.
Smaller pockets of contaminants (ie Howes and Potter’s Favorite) were not noticed in the visual inspection.
In the BG bed, no contaminants were noticed visually, as BG48 is very similar to BG. The three unknowns were also not able to be discerned visually.
In all, visual analysis was not perfectly accurate, but it was a useful indicator of genetic contamination, especially for larger blocks of contamination. It is suggested that visual analysis continue to be a guide when choosing samples to fingerprint genetically.
The decision to renovate the beds of the WCRS station in two phases was ultimately governed by location, elevation, water routing as well as yield from 2013-2017 and genetic contamination percent. The work done by Dr. Zalapa’s lab to enable this decisionmaking is thoroughly appreciated by the WCRS board of directors, and by all growers who benefit from research undertaken at the research station.
Genetic Fingerprinting at the Wisconsin Cranberry Research Station: Part 1, featuring cultivars found in existing WCRS beds, appears in the Cranberry Crop Management Journal vol. 35 issue 2.
Daniel Matusinec, Andrew Maule, Eric Wiesman, Amaya Atucha, Mura Jyostna Devi & Juan Zalapa (2022) The New Cranberry Wisconsin Research Station: Renovation Priorities of a ‘Stevens’ Cranberry Marsh Based on Visual Mapping, Genetic Testing, and Yield Data, International Journal of Fruit Science, 22:1, 121-132, DOI: 10.1080/15538362.2021.2014016 https://www.tandfonline.com/doi/full/10.1080/15538362.2021.2014016
This article was posted in Cranberry and tagged Allison Jonjak, Cranberries, genetic fingerprinting, Juan Zalapa, WCRS, WI Cranberry Research Station, Wisconsin Cranberry Research Station.
